
Using SIFT and PolyPhen to Predict Loss-of-Function and Gain-of-Function Mutations | Genetic Testing and Molecular Biomarkers
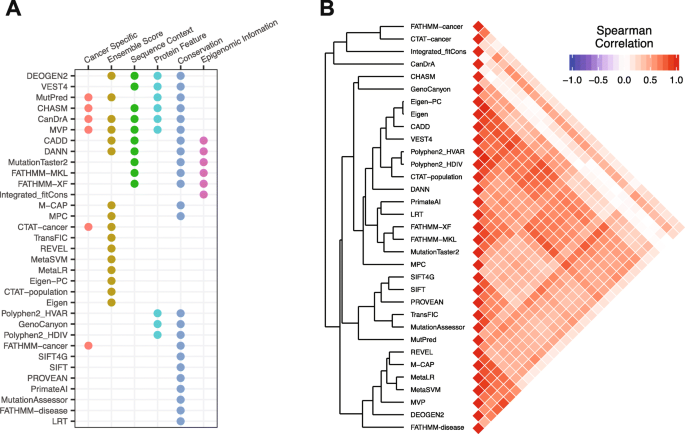
Comprehensive assessment of computational algorithms in predicting cancer driver mutations | Genome Biology | Full Text

Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm | Nature Protocols
Prediction of the Damage-Associated Non-Synonymous Single Nucleotide Polymorphisms in the Human MC1R Gene | PLOS ONE

PDF) Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm | prateek kumar - Academia.edu
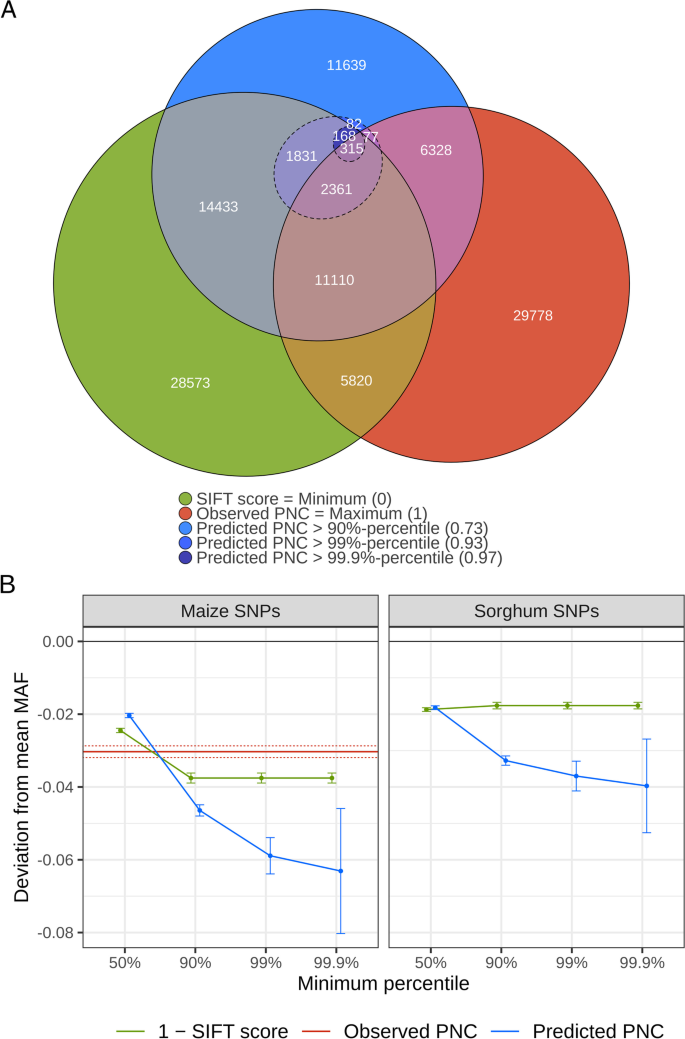
Prediction of evolutionary constraint by genomic annotations improves functional prioritization of genomic variants in maize | Genome Biology | Full Text
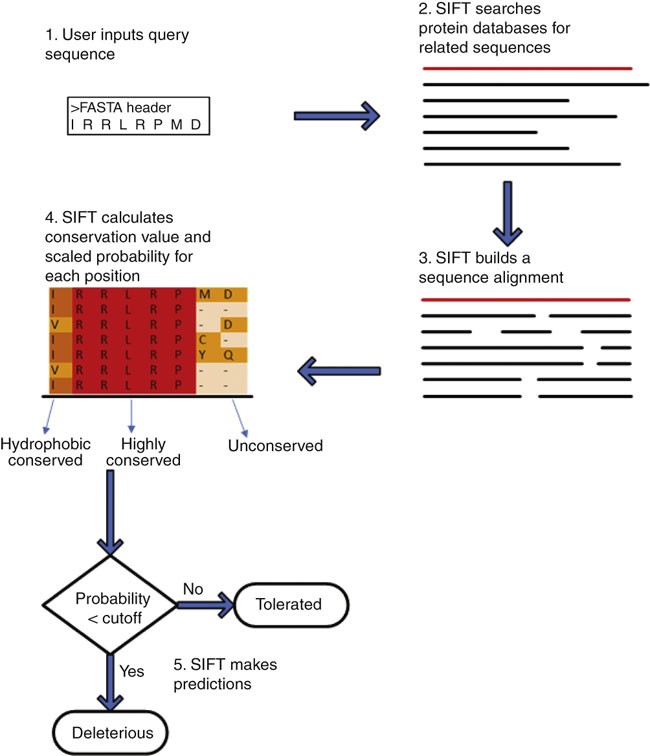
Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm | Nature Protocols
Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm - Document - Gale OneFile: Health and Medicine
Identification of Deleterious Single Nucleotide Polymorphism (SNP)s in the Human TBX5 Gene & Prediction of Their Structural

![PDF] SIFT web server: predicting effects of amino acid substitutions on proteins | Semantic Scholar PDF] SIFT web server: predicting effects of amino acid substitutions on proteins | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/051f851112b1ace45c86a4d37f0478a3e5d2fa6d/3-Figure1-1.png)







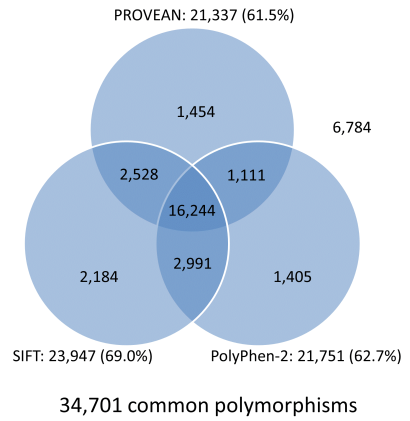

![PDF] SIFT: predicting amino acid changes that affect protein function | Semantic Scholar PDF] SIFT: predicting amino acid changes that affect protein function | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/23ca99cc5b24ff959e195879413fdf66f26e8373/3-Figure2-1.png)
